Mutational signature in colorectal cancer caused by genotoxic pks + E. coli
Mutational signature in colorectal cancer caused by genotoxic pks + E. coli
Cayetano Pleguezuelos-Manzano*, Jens Puschhof*, Axel Rosendahl Huber*, Arne van Hoeck, Henry M Wood, Jason Nomburg, Carino Gurjao, Freek Manders, Guillaume Dalmasso, Paul B Stege, Fernanda L Paganelli, Maarten H Geurts, Joep Beumer, Tomohiro Mizutani , Yi Miao, Reinier van der Linden, Stefan van der Elst, Genomics England Research Consortium; K Christopher Garcia, Janetta Top, Rob J L Willems, Marios Giannakis, Richard Bonnet, Phil Quirke, Matthew Meyerson, Edwin Cuppen, Ruben van Boxtel#, Hans Clevers#Nature 2020 Apr;580(7802):269-273. doi: 10.1038/s41586-020-2080-8. Epub 2020 Feb 27.
Abstract
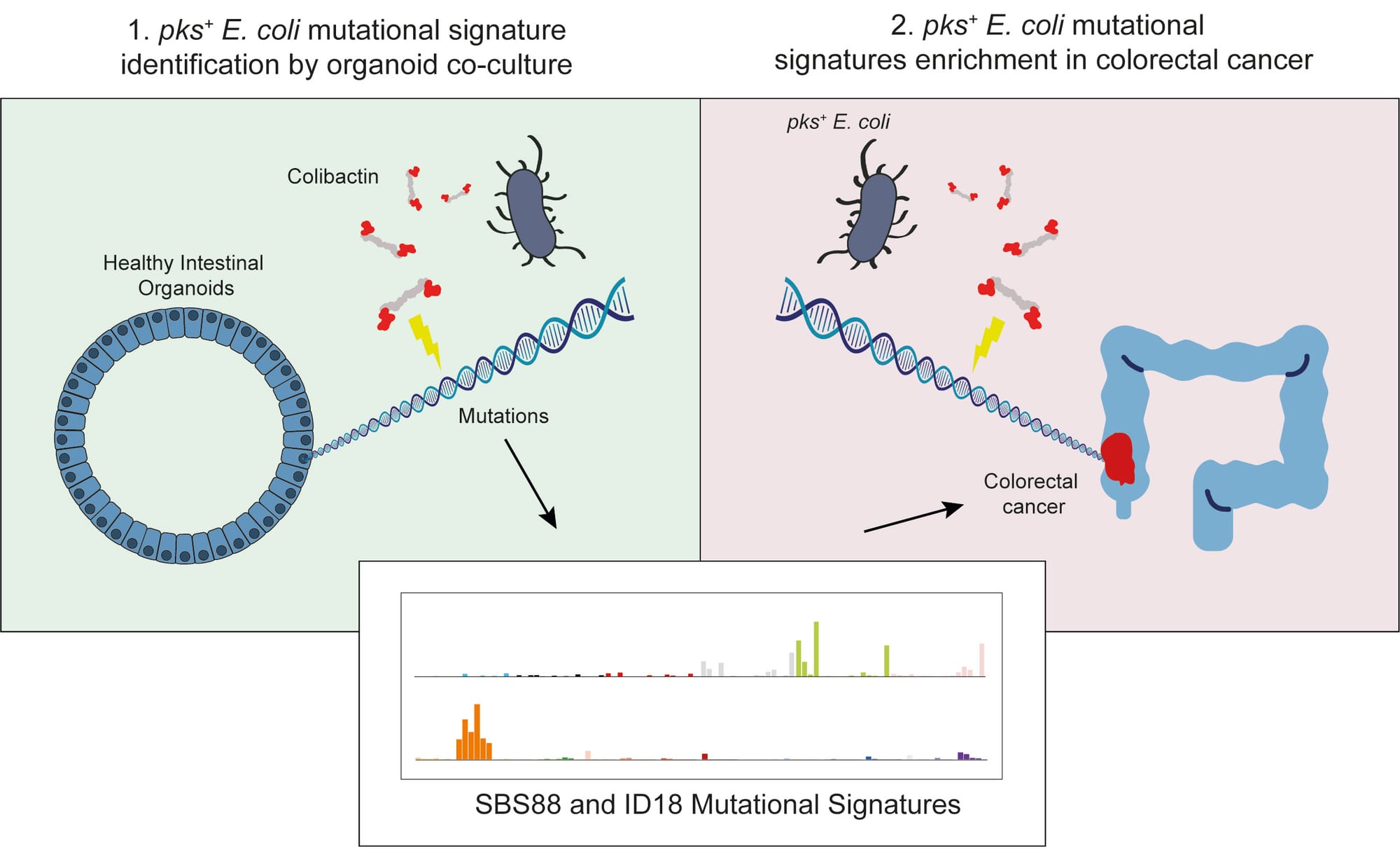
Various species of the intestinal microbiota have been associated with the development of colorectal cancer1,2, but it has not been demonstrated that bacteria have a direct role in the occurrence of oncogenic mutations. Escherichia coli can carry the pathogenicity island pks, which encodes a set of enzymes that synthesize colibactin3. This compound is believed to alkylate DNA on adenine residues4,5 and induces double-strand breaks in cultured cells3. Here we expose human intestinal organoids to genotoxic pks+ E. coli by repeated luminal injection over five months. Whole-genome sequencing of clonal organoids before and after this exposure revealed a distinct mutational signature that was absent from organoids injected with isogenic pks-mutant bacteria. The same mutational signature was detected in a subset of 5,876 human cancer genomes from two independent cohorts, predominantly in colorectal cancer. Our study describes a distinct mutational signature in colorectal cancer and implies that the underlying mutational process results directly from past exposure to bacteria carrying the colibactin-producing pks pathogenicity island.


